Traceability
Example of genomic analysis for traceability - Trout of the Orb
The Orb (Hérault) Mediterranean basin, a river in the south of France with a surface area of 1580 km2, has been supplemented with domestic trout (Salmo trutta) of Atlantic and Mediterranean origin since the end of the 1960s. The objective is to increase the local Mediterranean populations for recreational fishing.
In the study by Leitwein et al. (2018), the use of 86,175 SNPs (single nucleotide polymorphisms) markers from RAD sequencing, allowed to identify the proportion of domestic (Atlantic and Mediterranean) alleles in wild populations of the Orb basin.
This unable the identification of domestic Atlantic individuals (i.e. escaped from fish farms) in the Gravezon river, individuals of Mediterranean domestic origin in the Mare river, as well as hybrid individuals with Atlantic and Mediterranean domestic alleles in the three wild rivers (Figure 1).
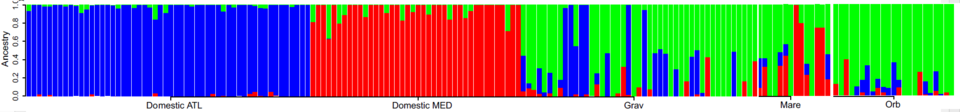
Figure 1 from Leitwein et al. 2018, Molecular Ecology.
Each bar represents the proportion of Atlantic domestic ancestry in blue, Mediterranean domestic in red and wild in green for the three populations of Gravezon (Grav), Mare (Mare) and Orb (Orb).
Studying the distribution of Atlantic and Mediterranean domestic haplotypes (segments) allowed to estimate the age of supplementation for each of the three rivers.
The study show that Atlantic domestic individuals were introduced on average 24.01 generations ago in the Gravezon river, 22.31 generations in the Mare river and 30.9 generations ago in the Orb river (Figure 2).
Similarly, the time of domestic Mediterranean trout introduction was estimated at 11.51 generations in the Gravezon River, 25.07 generations in the Mare River and 23.23 in the Orb River (Figure 2).
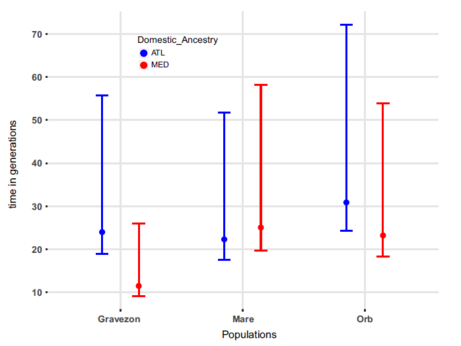
Figure 2 from Leitwein et al. 2018, Molecular Ecology.
Time in number of generations since the introduction of Atlantic domestic (blue) and Mediterranean domestic (red) individuals in the three populations Gravezon, La Mare and Orb.
Development of genomic tools - The high density trout chip
As a result of this study, we developed a high-density chip of 12,204 SNPs markers distributed homogeneously along the 40 linkage groups (LG) of the trout (Salmo Trutta) (Figure 3 ; Saint-Pé et al. 2019).
This chip allows for low-cost, consistent, and comparable management between different population genetic studies.
The chip is both effective in distinguishing trout of Atlantic and Mediterranean origins as well as hybridization levels.
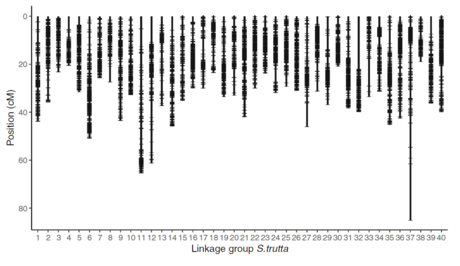
Figure 3 from Saint-Pé et al (2019), BMC genomics.
Positions (in centiMorgan) of the 12,204 SNPs markers along the 40 linkage groups of the common trout (Salmo trutta).
Stick pump
- Four filters in parallel
- Pumping speed: 40 L/min
- Integrated flow meter
- Makita 18V battery not included
- 20 µm nylon filter, 47 mm diameter


Yellow Submarine
- Three parallel filtration systems
- Filterd volume per vacuum depends on speed (approx. 50L/filters in 10 min at 6 knots
- 20µM nylon filter, 47 mm diameter
- Towed behind boat


GENEXPERT SAS
SIREN 952193837 - RCS Montpellier - NAF 7112B
