Conservation
Genetic diversity of trout in the Orb River
The use of 196K SNPs markers allowed us to estimate the genetic diversity and the inbreeding index of the common (Salmo trutta), wild (La Mare, Gravezon and Orb) and domestic (Atlantic and Mediterranean) trout populations of the Orb basin.
We were able to highlight that the wild Gravezon population had lower genetic diversity and high inbreeding rates compared to the wild La Mare and L'orb populations (Table 1; Leitwein et al. 2016).
Furthermore, we demonstrated that the Mediterranean domestic strain was very low in diversity and highly inbred in contrast to the Atlantic domestic strain, which had higher genetic diversity.
These results demonstrate that the Mediterranean domestic strain requires special management to increase genetic diversity.
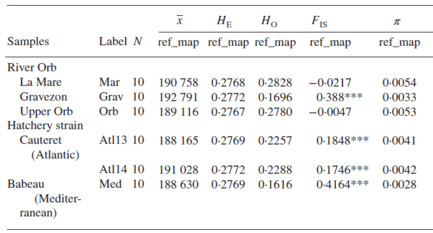
Table 1 from Leitwein et al. 2016, Journal of Fish Biology.
Summarize for each wild river in the Orb basin (La Mare, Gravezon, and Orb) and for the two domestic strains (Atlantic and Mediterranean), of (N) number of individuals, (x) average number of SNPs markers, (He) expected heterozygosity, (Ho) observed heterozygosity, (Fis) inbreeding index, and (π) nucleotide diversity. ***significant value p<0.0001
Genomic landscape of domestic ancestry in brook trout
The brook trout (Salvelinus fontinalis) is a species of high socio-economic interest in Canada. In the province of Quebec, a domesticated strain was created about 100 years ago in order to repopulate the rivers for sport fishing. Each year more than 650 tons of brook trout are released into Canadian rivers, promoting hybridization events between domestic and wild individuals (Leitwein et al. 2019).
Using 11,803 SNP markers, the ancestry of individuals could be inferred and the proportion of domestic alleles in wild rivers estimated. Using the linkage information and the proportion of domestic ancestry, it was possible to identify the hybrid classes of individuals. Individuals were categorized as either (i) "recent hybrids" i.e., offspring of early generations of interbreeding between wild and domestic individuals (<10 generations); or (ii) "ancient hybrids" i.e., offspring from distant mixing (>10 generations). Once individuals were assigned to their hybrid class, the rate of introgression of domestic alleles along the genome could be estimated. This approach revealed high variability, with genomic regions of high and low domestic ancestry, which are likely to be under natural selection (Figure 1; Leitwein et al. 2019).
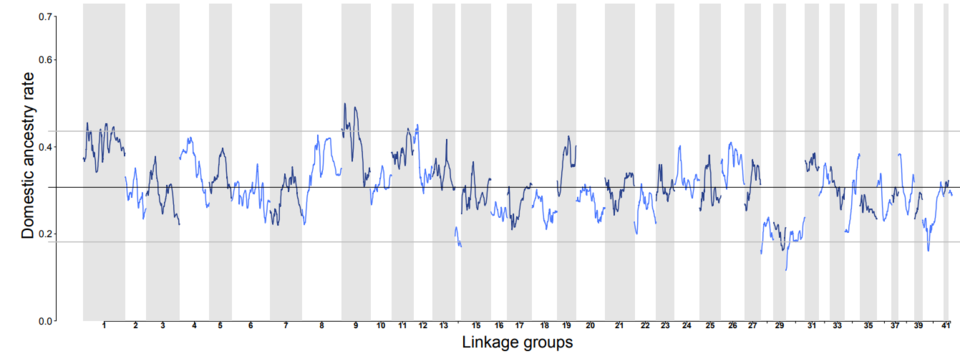
Figure 1 from Leitwein et al. 2019. Molecular Ecology.
Domestic ancestry rates along the 41 brook trout linkage groups (LG) for individuals in the recent hybrid category.
Effects of introducing domestic alleles
In the case of brook trout, we were able to observe variations of domestic ancestry in wild populations. For hybrid individuals, beneficial effects of the introgression of domestic alleles in wild populations were mainly observed. The domestic alleles bring (i) diversity in regions of the "wild" genome with low genetic diversity and (ii) potentially deleterious recessive alleles.
However, beyond these potential beneficial effects, purging effects of domesticated alleles have also been observed. These mechanisms reflect negative effects of introducing domestic alleles into certain regions of the genome (Leitwein et al. 2019 et 2021).
For example, on linkage group 26 (Figure 2) we can observe an excess of domestic ancestry associated with potentially domestic recessive alleles.
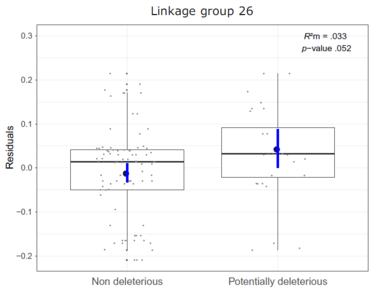
Figure 2 from Leitwein et al. 2019 Molecular Ecology.
Excess or deficit of domestic ancestry as a function of the presence of non-deleterious or potentially deleterious alleles for linkage group 26 in ancient hybrid brook trout individuals.
Stick pump
- Four filters in parallel
- Pumping speed: 40 L/min
- Integrated flow meter
- Makita 18V battery not included
- 20 µm nylon filter, 47 mm diameter


Yellow Submarine
- Three parallel filtration systems
- Filterd volume per vacuum depends on speed (approx. 50L/filters in 10 min at 6 knots
- 20µM nylon filter, 47 mm diameter
- Towed behind boat


GENEXPERT SAS
SIREN 952193837 - RCS Montpellier - NAF 7112B
